The platform is open to regular or punctual users from the INM and the exterior (researchers, post-docs, doctoral students, students and technical staff) who wish to carry out in situ hybridization, real time PCR, Nanostring analysis or sequencing.
The manager of the gene expression platform is responsible for:
- organization and management of the platform
- coordination of the different users
- advice, direction and training for future users
- development and adaptation of methodology to specific needs
1. In Situ hybridization
Technical and scientific referent: Stéphanie Ventéo
Users have all the necessary equipment (incubators, waterbaths, micropipettes, hybridisation oven, fume hood, microscopes to follow the colour development) and consumables at their disposition for carrying out in situ hybridization on whole tissues or section.
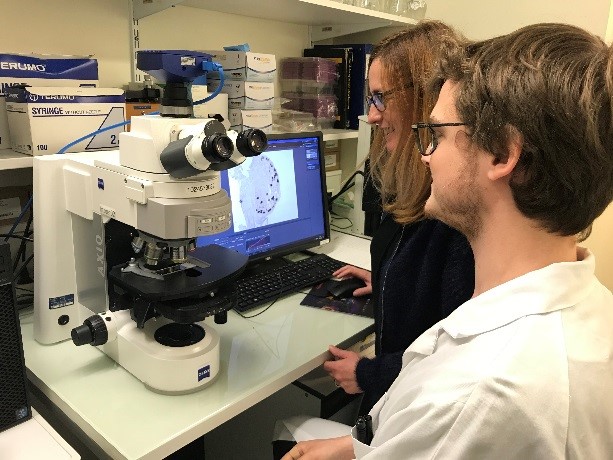


The in situ hybridisation laboratory is an RNase-free environment in which users have all the material necessar for extraction of RNA (Polytron, Ultratorax, centrifuges).
-
- simple in situ hybridization
- 2-colour double in situ hybridization
- hybridization in situ coupled with immunhistochemistry
- microRNA in situ hybridization
- fluorescent double in situ hybridization
EXAMPLES OF LABELING ON DIFFERENT TISSUES STUDIED AT THE INM


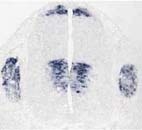
Retina Cochlea Spinal cord and Dorsal Root Ganglions

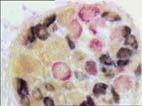
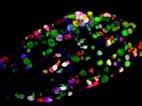
Nerve Double in situ hybridization ISH coupled with immunhistochemistry
Education

National training provided as part of the Continuing Education of INSERM .
The next session is programmed for 2021.
Leurs N et al., Frontiers in Genetics 12, 620659, 2021
Milman A et al., J Physiol. doi: 10.1113/JP281044, 2021
Ventéo S et al., Cell Rep., 29:2953-2960, 2019
Debiais-Thibaud M et al, Mol Biol Evol., 36:2265-2276, 2019
Coque E et al., Proc Natl Acad Sci USA pii: 201815961, 2018
Enault S et al., BMC Evol Biol. 8:127, 2018
Rivat C et al., Nat Commun. 9:1042, 2018
Ventéo S et al., Sci Rep. 6:36407, 2016
Ohayon D et al., Cell Rep. 17:1473-1481, 2016
Bouilloux F et al., Elife pii: e11627, 2016
2. Real time PCR
Technical referent: Stéphanie Ventéo
Scientific referent: Ilana Méchaly
Users have all the necessary equipment at their disposition for carrying out real time PCR:
Light Cycler 2.0 32 capillaries Light Cycler 480 plates 96 wells


The DNA free laboratory is an DNA-free environment in which users can do all the mix for PCR without the DNA matrix.
3. Nanostring Sprint Profiler
Technical referent: Cécile Monzo
Scientific referent: Carole Crozet
The Ncounter Sprint Profiler from Nanostring (https://www.nanostring.com/products/ncounter-analysis-system/sprint-profiler/) allows digital analysis of multiple elements from the same sample (microRNA samples, Total RNAs, proteins, genomic DNA, from paraffin-embedded tissues). It offers the possibility to analyze panels of miRNA, mRNA (up to 800 targets analyzed in the same sample), fusion of genes, lncRNAs, proteins (up to 200 proteins) defined by Nanostring or custom (see with Nanostring for the design).
It is also possible to study numerous pathways in a single experiment and in many fields (immunology, neurosciences, oncology, infectious diseases, clinical research …).
https://www.nanostring.com/products/ncounter-analysis-system/ncounter-systems-overview/
4. SeqStudio Genetic Analyzer (ThermoFisher)
Technical referent: Grégor Dubois
Scientific referent: Gaël Manes
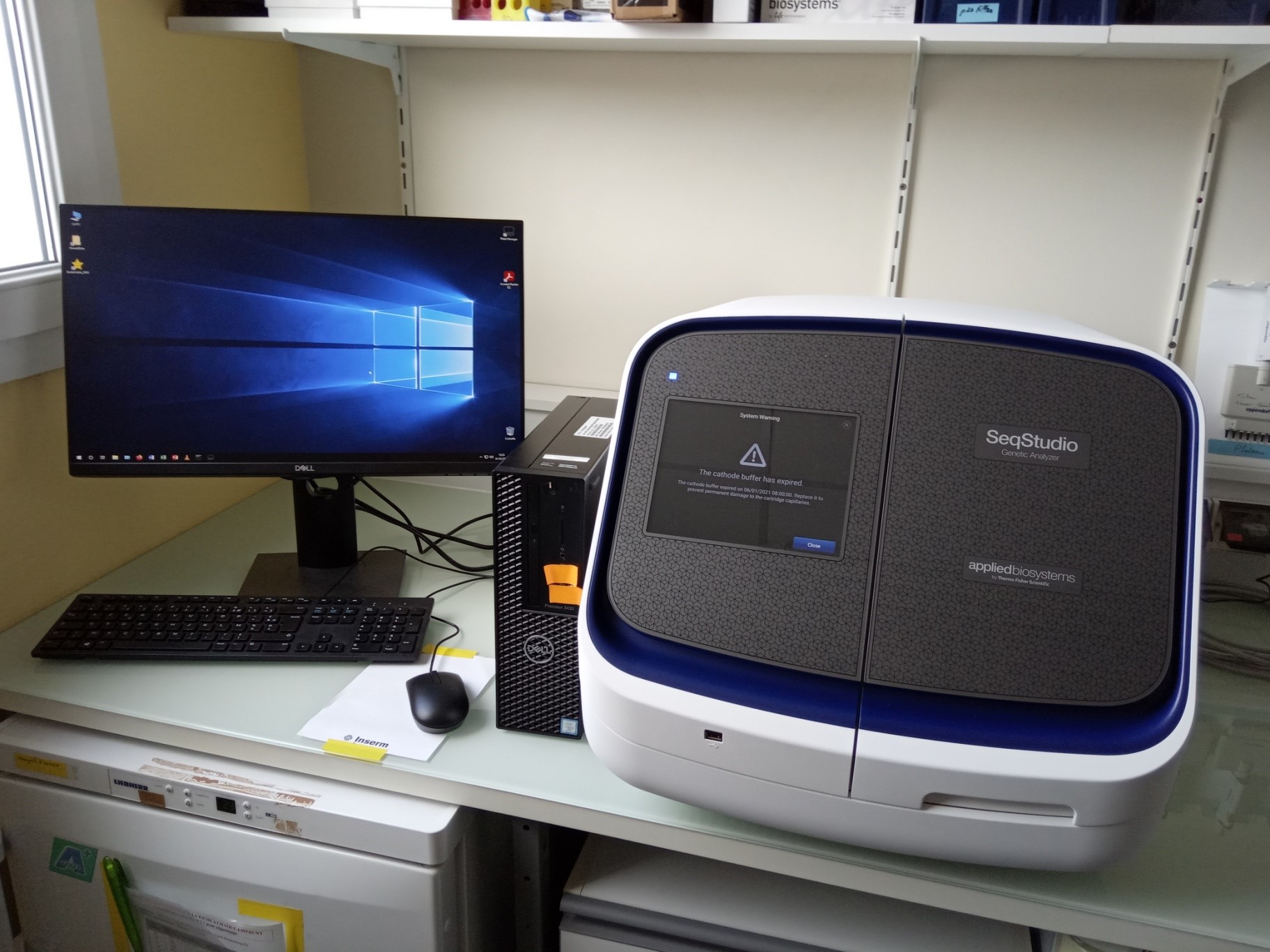

The ThermoFisher SeqStudio sequencer is a four-capillary, compact electrophoresis system operating with all-in-one reagent cartridges. Its use is simple and intuitive.
The SeqStudio can be used for different applications such as:
- Sanger sequencing (mutation detection, NGS variant confirmation)
- Plasmid sequencing
- Human cell lines authentication
- CRISPR-cas9 genome editing analysis
- Genotyping of fluorescence-coupled fragments
Sequencing analysis and GeneMapper software enable rapid analysis of sequencing and genotyping results. The sequencing facility has a thermocycler and a plate centrifuge dedicated to sequencing reactions.
Contact
















